While SB2.0’s headline goal is improved Sauvignon Blanc clones, NZW’s foundational genetics programme has already created long-lasting assets for the industry: new disease and virus testing capability, scalable screening tools, modern genetic “fingerprinting”, robust data systems, and stronger global and local partnerships. Programme Manager Dr Darrell Lizamore reflects on what’s now in place, and why it matters for winegrowers.
One of the most gratifying parts of my work has been seeing the enthusiastic response from visitors to our new breeding vineyard. With the vine population expected to reach 10,000 vines this season, it’s a tangible (and large) example of the progress we’ve made toward developing and selecting new clones of our dominant national variety.
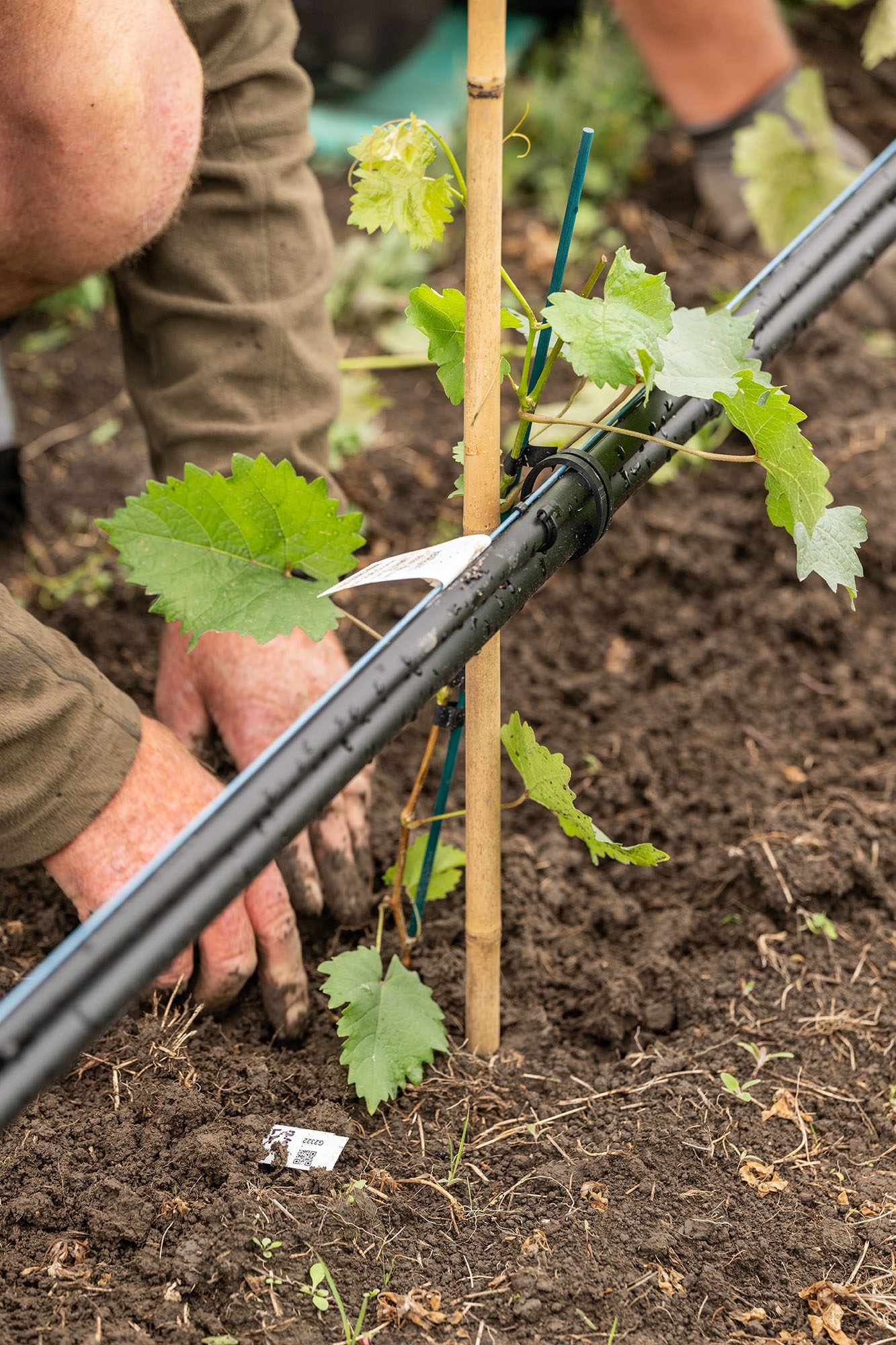 Less visible are the modern resources for grapevine improvement we’ve built in the background. Developing new vines is a long game, but the capability to produce and screen them faster, more reliably, and with less risk will accelerate progress and reduce costs for years to come. This article will shed some light on that capability and how it’s already being used beyond the Sauvignon Blanc 2.0 Programme (SB2.0).
Less visible are the modern resources for grapevine improvement we’ve built in the background. Developing new vines is a long game, but the capability to produce and screen them faster, more reliably, and with less risk will accelerate progress and reduce costs for years to come. This article will shed some light on that capability and how it’s already being used beyond the Sauvignon Blanc 2.0 Programme (SB2.0).
The right people and the right place
One of the early challenges for our new programme was building a team with the vision, skills, and motivation to tackle it effectively. We needed scientists and collaborators, industry experts and technical advisors, investor representatives and people with programme governance experience, and engagement and communications specialists. With limited resources compared to other breeding programmes, it felt even more critical to get this right from the outset. Fortunately, we’ve been able to partner with talented and generous people in New Zealand and overseas who see the value in what we’re trying to achieve.
We’ve brought together a passionate team with expertise in genetics, plant physiology, bioinformatics, molecular biology and viticulture. They’ve rapidly grasped the needs of our wine industry through BRI’s relationships, closing the gap between real-world priorities and research innovation.
Improving our ability to measure
Plant breeding depends on producing large populations of diversity and then finding outstanding individuals among them. Whether the criteria are vine health, physical traits, or phenological data, there is ‘noise’ introduced by subjective observation and environmental variation that needs to be managed. The less reliable the measurements are, the more inefficient selection becomes, wasting resources and delaying progress.
Over the past year, SB2.0 has invested in optimising approaches for measuring the traits central to our Programme, including new methods, equipment, training, and data systems. These are the parts of research that typically go unnoticed when they work well, but that make the difference between a confident result and a lucky guess. What’s more, this is a resource the industry can use to take on future challenges and opportunities, from reducing inputs to adapting to a changing climate.
Stronger diagnostics
We implemented molecular diagnostics (PCR and amplicon sequencing) to confirm the absence of phylloxera in the new breeding vineyard. These tools allow us to detect pathogens from soil samples with high sensitivity. We also developed in-house protocols for leafroll virus detection so that we can test and receive OddVines (bud sports identified by commercial growers) throughout the year. Growers can now report OddVines in Vure, making it easier to flag vines during normal vineyard work.
A modern vine ID system
In 2023, we completed a Sauvignon Blanc reference genome, providing a baseline for identifying new genetic changes. Since then, the team has characterised differences among commercial Sauvignon Blanc clones and about 100 of the new clones produced in this programme. This means we have a greater understanding about what types of genetic changes occur between clones and the traits they affect, and a way to easily tell one clone from another by DNA testing.
Our genome assembly pipeline has also been used to generate high-quality references for varieties such as Chenin Blanc, Chardonnay, and Pinot noir, as well as rootstock varieties, helping enable wider grapevine breeding in the future.
To streamline and automate genetic testing, we developed new methods for DNA extraction and sequencing. These reduce our per-vine costs and can detect a wider range of genetic differences. We built bioinformatics pipelines that can identify clonal variation reliably, and can even map the molecular switches that control which traits are expressed. The data is kept in a database with digital field books, barcode integration, and QR-code identifiers, and the software now runs on national high-performance computing infrastructure, enabling stable production workflows.
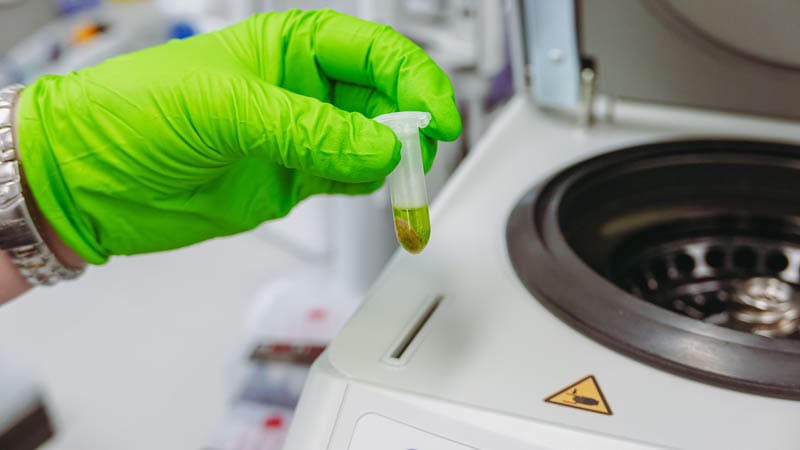 Part of the DNA extraction method
Part of the DNA extraction method
These tools are now being used to bring our national vine collection database into the 21st century and to characterise the impact of terroir and environmental stress on grapevine trait expression.
Making selection scalable
Powdery mildew screening is the clearest example of how we are attempting to scale trait selection. We standardised inoculum production and refined detached-leaf assays, then paired them with automation. The Blackbird imaging platform (developed by USDA/Cornell) is now being used in our lab locally to digitise mildew growth, using AI to distinguish mildew from leaf hairs. This turns large image datasets into consistent, objective scores in hours rather than weeks.
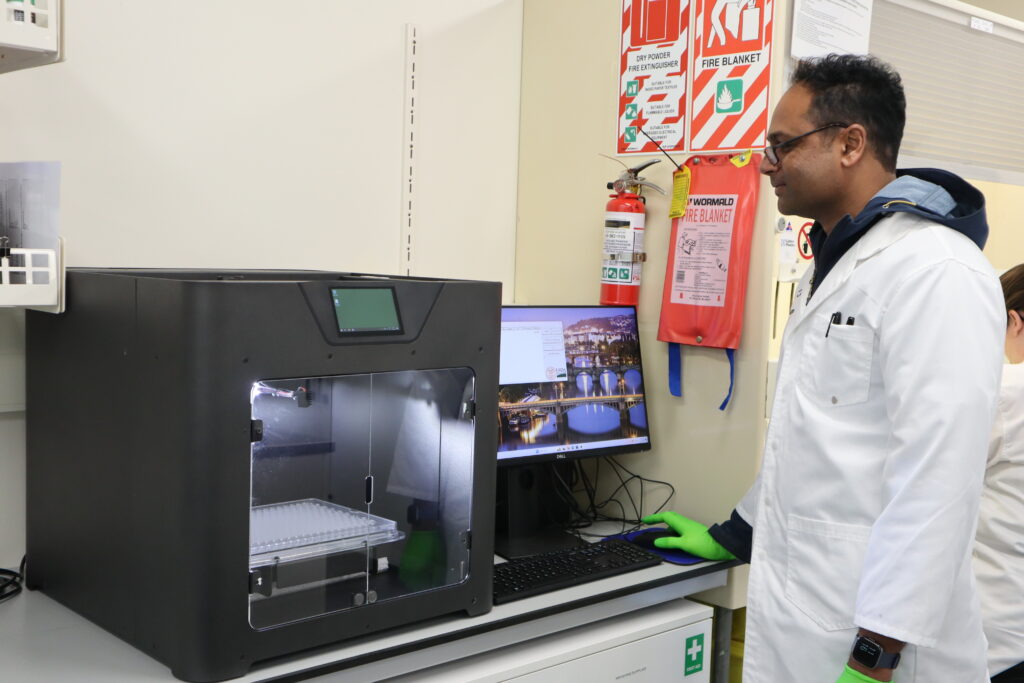 Research scientist Dr Bhanupratap Vanga with the Blackbird robot
Research scientist Dr Bhanupratap Vanga with the Blackbird robot
This new testing resource has been applied to fungicide resistance testing and is being used to survey fungicide resistance in local powdery mildew isolates and develop rapid diagnostics.
Measuring climate resilience
For drought and water-use efficiency, we ran a glasshouse pilot study with 80 vines to find indicators that are both informative and scalable. A collaboration with Bordeaux Sciences Agro (France) has provided new tools and experimental designs for rapid measurement of photosynthesis and transpiration.
For frost tolerance screening, we’re combining controlled lab experiments with field exposure, including adapting robotic imaging approaches, to detect bud injury. This work has been guided by leading cold-climate researcher Markus Keller at Washington State University.
From lab bench to vineyard row
After four years of sterile plantlet production, we can now take new clones from the lab, via the nursery, to the breeding vineyard with a greater than 97% survival rate. The vines then follow a special management programme to accelerate their growth through a juvenile phase to mature vines ready for screening to begin the following year.
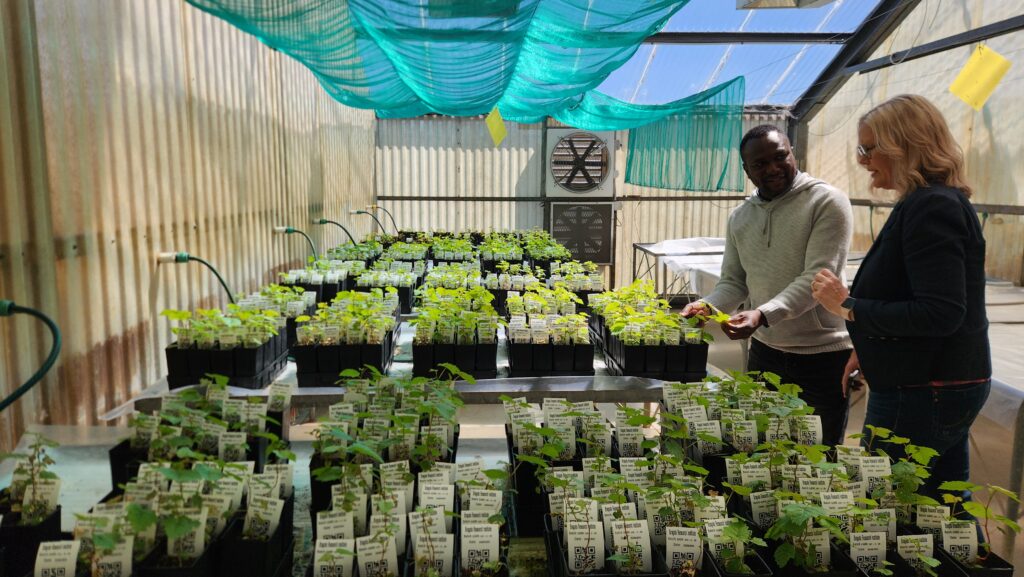 SB2.0 Clones in the nursery
SB2.0 Clones in the nursery
We’re now also testing ways to speed up their progress from early selection to commercial trials. An experiment in Marlborough this season will compare propagation strategies, including top-grafting buds from new clones onto established vines.
New gene technologies
This year, the government proposed new legislation to update gene technology definitions. BRI’s science team have provided technical expertise to support education and constructive discussion about gene technologies and market-access considerations. This has been met with enthusiastic industry engagement at Grape Days events, webinars, and with groups like Organic Winegrowers NZ.
Even if the industry chooses to move cautiously on new gene technologies, building capability in regulated research settings has value. We’ve tested the necessary steps for non-transgenic editing, including regeneration from grapevine protoplasts. These techniques would enable early validation of breeding strategies and may one day form the basis of new approaches for grapevine improvement, as demonstrated in overseas trials.
Networks and spin-offs
On an international scale, SB2.0 is a unique programme that has attracted interest and collaboration opportunities. We have forged direct connections with grape breeders in Europe and North America, including technical visits to UC Davis, Cornell, USDA, and E. & J. Gallo. We hosted breeders Valentin Blattner (Switzerland) and Peter Cousins (USA) to exchange knowledge on mildew resistance genetics, and we will host breeders from Geisenheim University and ICVV (Spain) in early 2026.
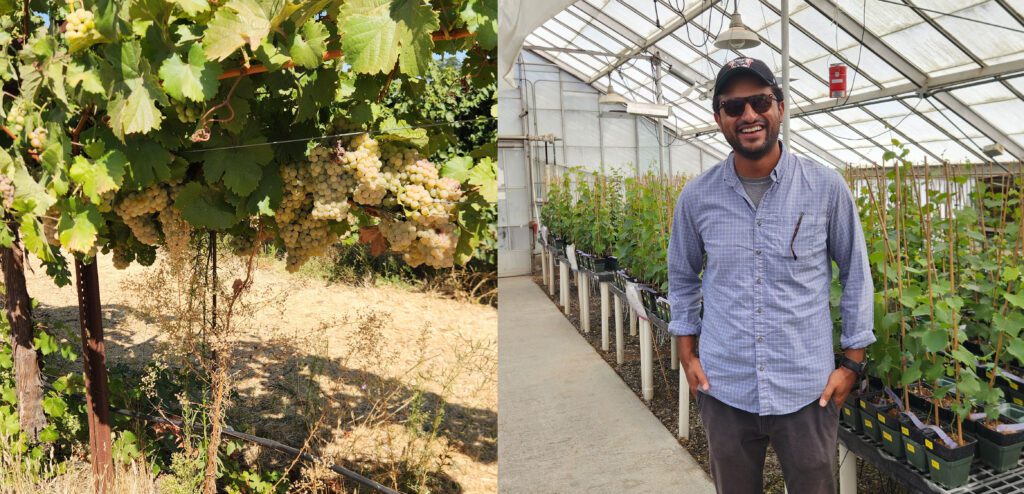 UC Davis breeding programme with breeder Luis Diaz Garcia
UC Davis breeding programme with breeder Luis Diaz Garcia
Locally, the platform connects grantor companies, regional grower communities, and industry committees with science through workshops, newsletters and in-person meetings. It improves two-way learning about what breeding can deliver and which improvements would be most impactful.
What comes next
Selection takes time, but the platform we’ve built uses modern technology and international connectedness to reduce the time and cost per decision. The next phase of SB2.0 is about selecting traits that make Sauvignon Blanc more resilient, such as mildew tolerance, drought response, and frost tolerance. This will mean running objective, repeatable screening workflows, connecting trait measures to genetics, and eventually moving the most promising material into pre-commercial trials.
Beyond the core goals, SB2.0 tools are already supporting spin-off projects, including fungicide resistance surveys, RNA-based treatments for leafroll virus, characterising the diversity of Pinot Noir clones, and breeding local disease-tolerant varieties together with the Bioeconomy Sciences Institute.
The programme offers more than the opportunity to develop the next Sauvignon Blanc clone. It’s also enhancing our ability to keep improving vines as climate, disease pressure, and markets evolve, using innovation and collaboration.
















