What is your leafroll number?
Arnaud G Blouin and Robin M MacDiarmid
The old frescos from Pompeii have been preserved for centuries by the ashes of the Mount Vesuvius eruption (79 AD). It is in some of these paintings that the oldest grapevine virus symptoms can be observed. Indeed, the distorted grapevine leaves depicted resemble those caused by the virus Grapevine fanleaf virus (fanleaf). Vegetative propagation of grapevines has been a common practice for the last 6000 years, resulting in humans being the main vector of grapevine viruses with additional transmission routes superfluous to the point that some cultivars, such as the Vitis vinifera Red globe, can be identified by the viruses they host (leafroll 2).
The missionary Samuel Marsden is often credited with having planted the first grapevine in New Zealand in 1819. With the current knowledge of the ubiquity of the grapevine viruses, we can probably also attribute to him the first grapevine viruses in New Zealand. Further evidence of the presence of virus disease in New Zealand grapevines can be traced to 1902; in the 10th report of the Department of Agriculture, Romeo Bragato described Cabernet Sauvignon vines that produced no fruit and were easily distinguished from fruit-bearing vines by the early reddening of their leaves. These descriptions could be attributed to the virus Grapevine leafroll-associated virus 3 (GLRaV-3 or leafroll 3). By the 1960s, such leafroll disease was known to be widespread. Subsequently, the virus has been recognised as the most detrimental in the country. In the first official report that specifically describes leafroll disease, McKissock also reported fanleaf, an important disease of vineyards at the time. The impact of fanleaf was assessed by Chamberlain in 1970, with high incidence reported in Auckland and Hawke’s Bay, the two main wine regions at the time. The alarming report by Mossop in 1986, relating a widespread occurrence of fanleaf and of the related virus Arabis mosaic virus (ArMV) in New Zealand, constitutes the last publication of viruses belonging to the genus Nepovirus in New Zealand grapevines. Ultimately, as a result of the absence of a vector in the country, and with a better control over planting material health, fanleaf, and other nepoviruses, are now considered eradicated from commercial vineyards. Since the mid-1980s to the beginning of this study, 17 other viruses have been reported in New Zealand grapevines.
Over the last four years, Arnaud Blouin has been undertaking his doctoral research titled “Virus diversity in New Zealand grapevines: Sequence, history and impact” under the supervision of Robin MacDiarmid at the University of Auckland and Plant and Food Research. The research that was funded and supported by the Rod Bonfiglioli Memorial Scholarship as well as Plant and Food Research has been reported previously in easy to read articles (Blouin AG, Bell VA, MacDiarmid RM 2016 The upsides of viruses. New Zealand Winegrower 97: 86-87, Anon. 2015 Sports and spots for survey. New Zealand Winegrower 89:57, Blouin AG, Ross H, MacDiarmid RM 2014 Virus diversity in New Zealand grapevines: sequence, ecology and impact. Overview of the Rod Bonfiglioli scholarship research project. New Zealand Winegrower 86:72-73, Massey E 2018. Exploring NZ’s vineyard virome. New Zealand Winegrower 108: 22-23). More detailed information about the research can be found within the seven research papers that comprise his thesis. Now that the thesis is submitted, we have updated its outcomes in this article.
Plants tested
The “virome” is defined by the totality of the viruses present in one environment, in this case the New Zealand environment. The first part of this study was to establish a reliable assay to detect all viruses present in a sample. The second part was to select which plants to test. We have collected 225 grapevines from different cultivars and regions that could be divided into four groups: First, the background virome of the commercial vineyard was assessed from 166 Sauvignon blanc samples collected from four vineyards within two major wine regions of New Zealand (labelled as the SB group). Secondly, the historical introduction of viruses into New Zealand was assessed from 19 plants collected from the New Zealand Winegrowers’ germplasm collection, the so-called “low-health” block (GC-LH group). The third source sampled 16 grapevines that had undergone a virus elimination process and were planted in the “high-health” block of the germplasm collection (GC-HH group). Lastly, potential virus-disease correlations were assessed from 24 commercial grapevine samples collected over three years by growers across New Zealand concerned about their symptomatic vines (DR group, Figure 1).
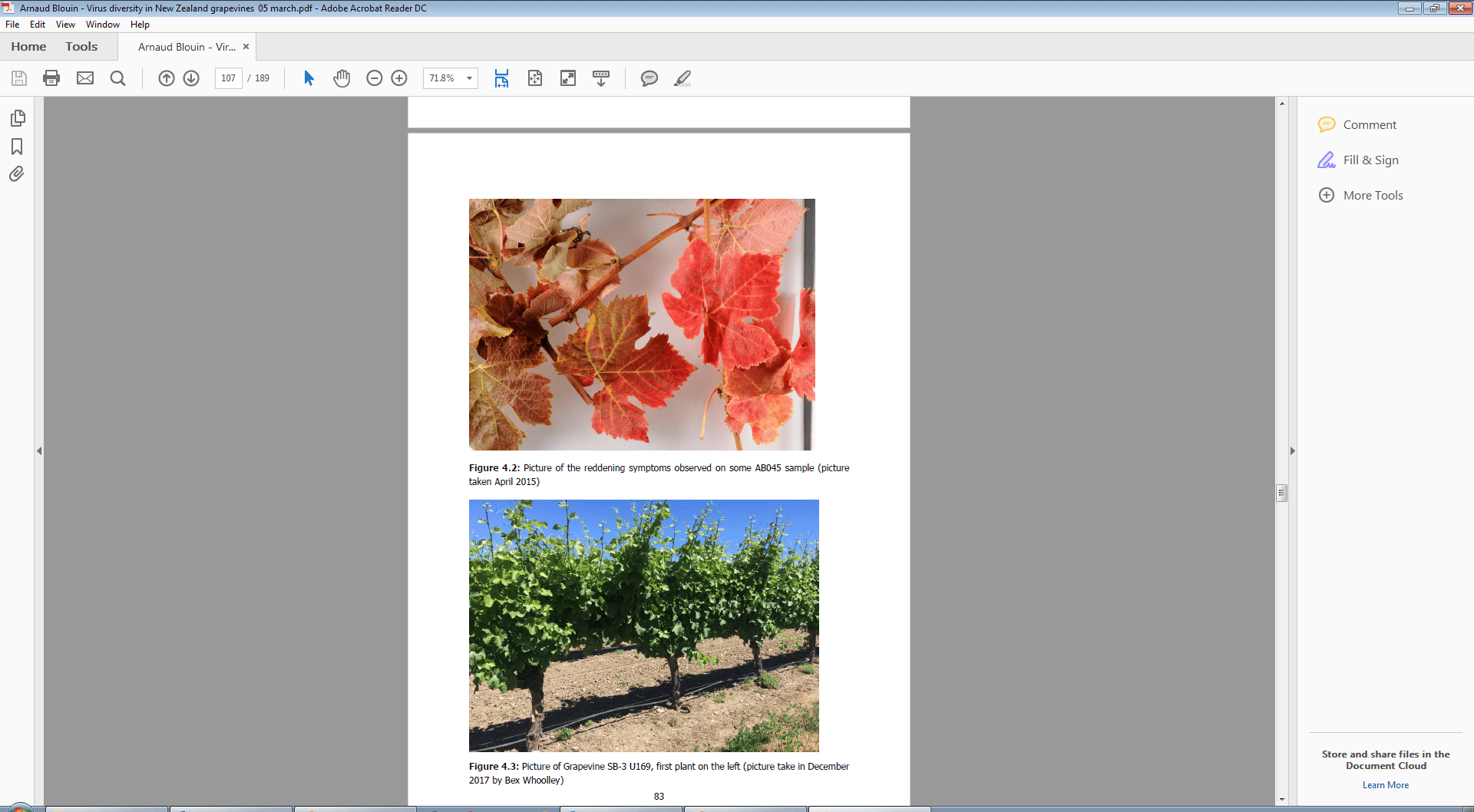
Figure 1. Picture of the reddening symptoms observed on Syrah sample AB045 infected with Grapevine rupestris stem pitting-associated virus (GRSPaV) and two common viroids but no candidate virus that might account for the symptoms. Picture taken by Arnaud Blouin (PFR) in April 2015.
Viruses detected
From the 17 viruses reported to be present in New Zealand at the start of this study, we have detected 14. In addition to fanleaf virus that was absent from New Zealand prior to this study, the three viruses not detected in this research belong to the Nepoviruses and include and ArMV, Tobacco ringspot virus (TRSV), and Tomato ringspot virus (ToRSV). These viruses are vectored through the soil by nematodes soil that are not present in New Zealand.
The 14 viruses already known to be present in the country are the leafroll viruses 1, 2, 3 and 4, the vitiviruses GVA, GVB and GVD, the ubiquitous Grapevine rupestris stem-pitting associated virus (GRSPaV) and five members of the family Tymoviridae that we group under the name ‘fleck-like viruses’; they are Grapevine fleck virus (GFkV), Grapevine rupestris vein feathering virus (GVFV), Grapevine red globe virus (GRGV), Grapevine asteroid mosaic-associated virus (GAMaV) and Grapevine Syrah virus-1 (GSyV-1).
In addition, we have detected three novel viruses belonging to the Vitivirus genus, the same genus as Grapevine virus A (GVA) and Grapevine virus B (GVB). Two of these viruses, Grapevine virus G (GVG) and Grapevine virus I (GVI) were not known to science before this research, the third one is novel and a distant relative to Grapevine virus E (GVE). Lastly, a virus named Grapevine geminivirus A (GGVA) was also detected during this study. This is the first report for of this virus in the country.
| n | GLRaV-1 | GLRaV-3 | GLRaV-4 | GLRaV-2 | GRVFV | GSyV-1 | GAMaV | GRGV | GFkV | GRSPaV | GVA | GVB | GVD | GVE-like | GVG | GVI | GGVA | |
| SB-1 | 44 | 0% | 2% | 0% | 7% | 14% | 20% | 0% | 9% | 7% | 32% | 0% | 0% | 0% | 0% | 0% | 0% | 0% |
| SB-2 | 43 | 0% | 0% | 0% | 2% | 7% | 7% | 0% | 5% | 7% | 100% | 0% | 0% | 0% | 0% | 0% | 0% | 0% |
| SB-3 | 40 | 0% | 5% | 0% | 0% | 0% | 60% | 0% | 8% | 0% | 0% | 0% | 0% | 0% | 3% | 3% | 0% | 0% |
| SB-4 | 39 | 0% | 0% | 0% | 0% | 0% | 85% | 0% | 0% | 0% | 0% | 0% | 0% | 0% | 0% | 0% | 0% | 0% |
| DR | 24 | 0% | 25% | 0% | 0% | 0% | 0% | 0% | 4% | 0% | 83% | 8% | 4% | 0% | 0% | 4% | 0% | 0% |
| GP-HH | 16 | 0% | 25% | 6% | 6% | 6% | 13% | 6% | 25% | 0% | 69% | 0% | 0% | 0% | 0% | 0% | 0% | 0% |
| GP-LH | 19 | 11% | 74% | 5% | 53% | 47% | 5% | 0% | 16% | 0% | 63% | 58% | 0% | 5% | 0% | 68% | 47% | 5% |
Table 1 Percentage of plants infected with one of the 17 viruses detected in the seven different categories: Sauvignon blanc vineyards in Hawke’s Bay (SB-1 and SB-2) or Marlborough (SB-3 and SB-4), mostly symptomatic samples received (DR), and samples collected from the germplasm high-health (GC-HH) and low-health (GC-LH) blocks. The number of samples per category is listed in column n.
Viruses distribution
As predicted, a high virus load was detected in the germplasm collection. Most of the vines deposited in this collection originated from the Te Kauwhata Viticultural Research Station, a national reference collection located in the Waikato region that was initially established in the late 1800s, then increased through imports of new grapevine cultivars. Over the ensuing decades, the cultivars were assessed and the best were distributed to New Zealand grape growers. Some of the accessions can be traced back to Romeo Bragato. The Te Kauwhata collection was not maintained in situ after the 1980s and was moved to different locations before being incorporated into the New Zealand Winegrowers germplasm collection in its current location in Lincoln. Lincoln is a region that is regarded as having low pressure from the insect vectors of viruses, in particular mealybugs. The plants are now self-rooted. Some of the plants, treated by thermotherapy to remove virus infections, were isolated in the GC-HH block alongside the most recent imports that have been screened through post-entry quarantine processes (1980s onward). From the 16 GC-HH plants sampled, an average of 1.6 viruses per plant were detected, as opposed to 4.6 virus infection per plant sampled in the GC-LH block (19 plants). Thus, the high-health and low-health terminology used to name those two blocks is justified.
The vitivirus GVG was discovered first in the New Zealand Winegrowers’ germplasm where it appears to be common (68% of the plants tested in the GC-LH). The virus was then detected in one plant within the Sauvignon blanc survey in Marlborough and another plant in a commercial vineyard in the Hawke’s Bay region. These single plant findings suggest that the virus was propagated outside the collection. A recent report including sequence data showed the same virus was detected in Croatia although there are substantial genetic differences to the one detected in New Zealand. The second new-to-science virus described is GVI, which is related to GVG within the vitivirus genus. GVI was detected in nine plants and these were all co-infected with GVG. This vitivirus GVI has not yet been detected outside the grapevine germplasm collection, and has not yet been reported outside New Zealand. A virus related to GVE was also detected in the survey of Sauvignon blanc from Marlborough (Figure 2). This was the first report of GVE (or a GVE-like virus) in the country. The plant was also co-infected with GVG. All these viruses belong to the genus Vitivirus with GVA, GVB and GVD that were also detected during this survey and GVF, GVH and GVJ reported in the same host overseas but not yet identified in New Zealand.
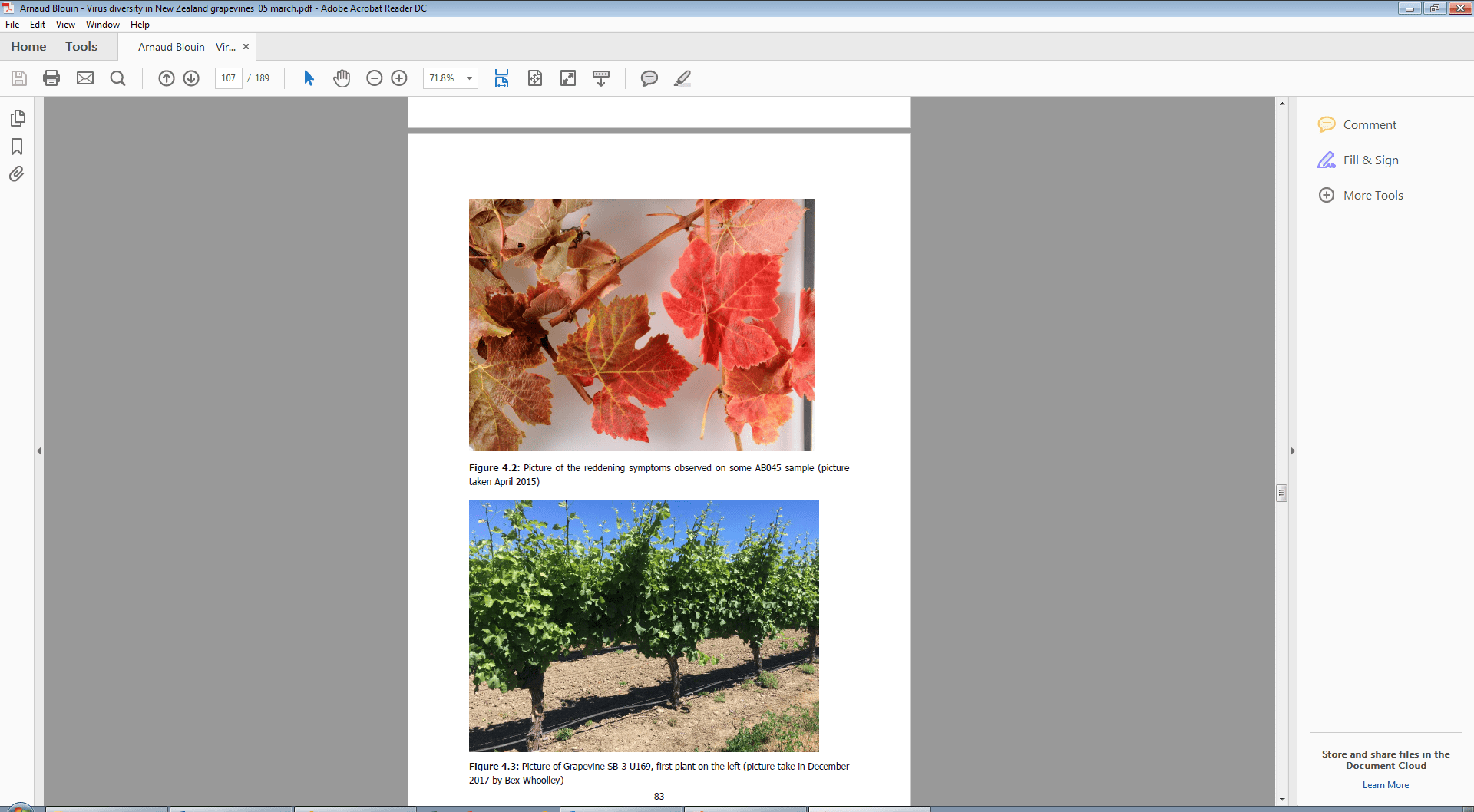
Figure 2. Picture of Grapevine SB-3 U169 (first plant on left) infected with the Grapevine vitivirus E (GVE), Grapevine vitivirus G (GVG), and leafroll 3. Picture taken by Bex Woolley (PFR) December 2017.
All the plants infected by one vitivirus in this research (GVA, GVB, GVD, GVE-like, GVG and GVI) were also infected with leafroll 3. In grapevine it is believed that the vitiviruses require the presence of a leafroll virus (GLRaV-1, -2, or -3) for its acquisition. However, a recent publication described that the vitivirus benefited from the presence of a leafroll virus to increase its replication in the Vitis host and therefore its chance of transmission rather than being dependent on transmission co-factors provided by the leafroll virus. The presence of the vitivirus does not seem to impact the leafroll virus concentration or transmission.
The detection of the geminivirus GGVA in one GC-LH grapevine constitutes the only finding of a DNA virus in the current study. This is the first report of the virus in New Zealand. The virus was originally described from the USA on imported vines from Korea. The imported plants displayed virus symptoms and were infected by multiple viruses. A subsequent survey of the US Department of Agriculture – Agricultural Research Services’ Clonal Germplasm Repository – found 15 additional GGVA infected plants with no correlation with symptoms. The virus was subsequently reported in Korea and China where it may be widespread, but no association with symptoms was established.
The single known plant positive for GGVA in New Zealand is an interspecific cross, Seibel 7052. The plant was also found to be infected with leafroll-2, GRSPaV, GAMaV, and a viroid, and no symptoms were observed at the time of collection (early 2016, Figure 3) or re-collection (January 2018). In the New Zealand grapevine variety register, the source and year of importation of this plant is absent but it was logged between TK00183 (Siebel 6339) and TK00188 (Siebel 10096) that were both imported in 1957 from the US Department of Agriculture. From the information available to date, we can conclude that GGVA in New Zealand is not causing severe symptoms and its spread is very limited; after possibly 60 years it has not spread to any of the other 15 plants sampled from the same germplasm collection. The impact and the spread of the virus therefore appears to be negligible. However, this finding should generate more interest in DNA viruses and their place in the New Zealand virome.
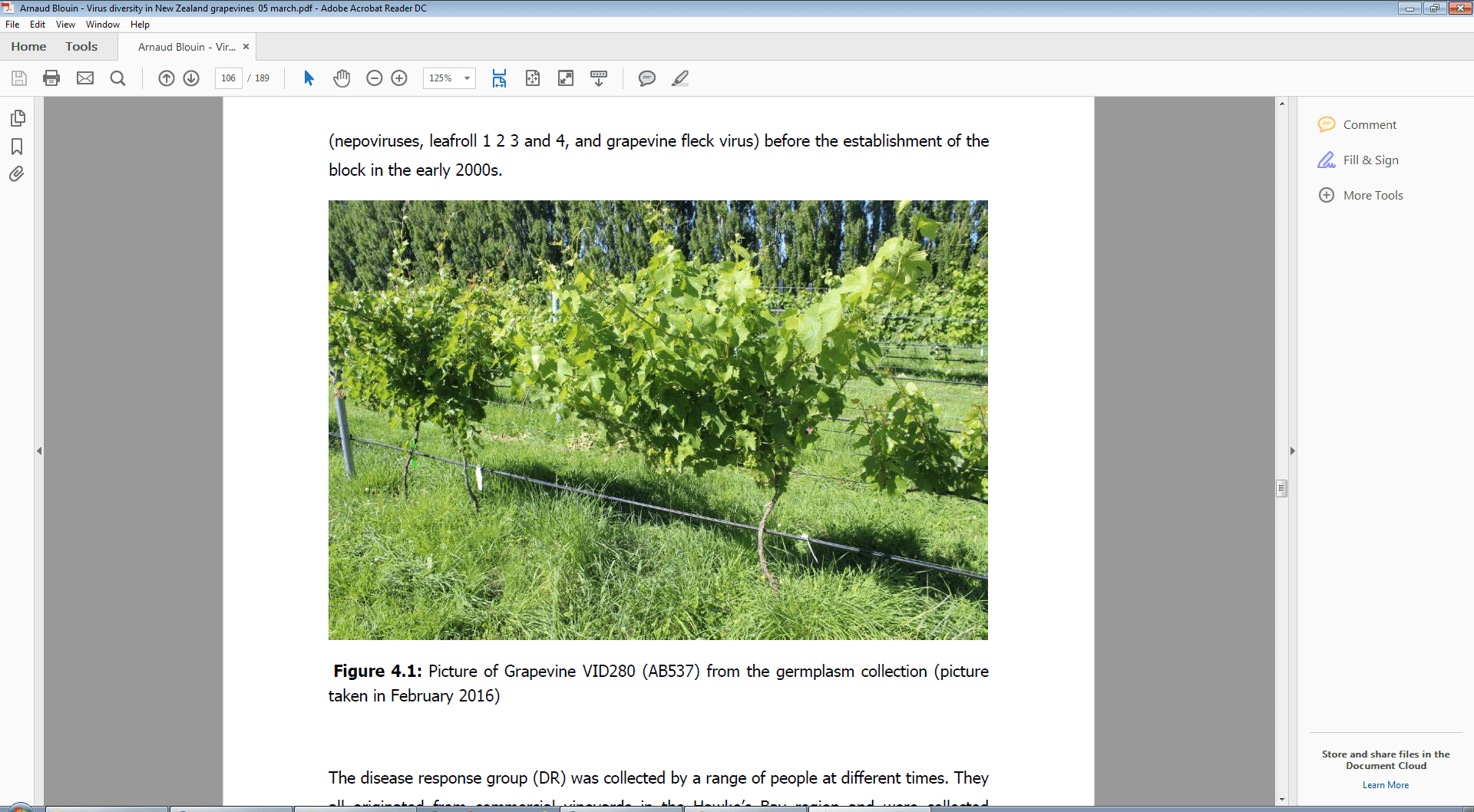
Figure 3. Picture of the Seibel 7052 grapevine VID280 (AB537) from the low-health germplasm collection (GC-LH) infected with Grapevine geminivirus A (GGVA), leafroll-2, Grapevine rupestris stem pitting-associated virus (GRSPaV), Grapevine asteroid mosaic-associated virus (GAMaV), and a viroid. Picture taken by Arnaud Blouin (PFR) in February 2016.
The grapevine survey in this current study comprised 225 plants and detected a total of 17 viruses. This study represents the largest survey worldwide for grapevine viruses and uses novel sequencing technologies that have identified the most viruses reported from a single study. Such a detection rate could insinuate that New Zealand vineyards are highly infected. However, closer examination demonstrates the exact opposite. The New Zealand commercial vineyards have a low virus incidence, with less than one virus detected per vine between the four Sauvignon blanc vineyards (average detection of 0.96 viruses per vine). In most cases, the viruses detected were GRSPaV or GSyV-1 and these are considered of low to no negative impact and may even be favourable (see below). Without these two prevalent viruses, the average number of viruses detected per plant drops to 0.2 per vine. The assortment of viruses that were detected at low incidence was surprising as seven additional viruses (leafroll-2, leafroll-3, GVE-like, GVG, GRVFV, GRGV and GFkV) were detected from the Sauvignon blanc vineyards. The relatively high health of the vineyards can be credited to the studious work of the nurseries to propagate clean material and the New Zealand Winegrowers’ Grafted Grapevine Standard (GGS) established and managed by the New Zealand Winegrowers that determines grapevine quality of plants sold by the nurseries, including the absence of GLRaV-3. This is also valuable information for nurseries, and it highlights the effective work of sanitation and risk awareness historically conveyed by scientists such as Dr Rod Bonfiglioli and Dr Richard Smart. It also reinforces the importance of using clean plant stocks in the nurseries to avoid spreading new pathogens.
The low virus incidence found in Sauvignon blanc surveyed in this study is also the result of strict import health regulations for Vitis since the Biosecurity Act in 1993. This legal document reformed the laws related to pest and unwanted organisms to New Zealand. The new viruses reported in this research were reported to the Ministry for Primary Industries (MPI) as reports of an infectious agent new to New Zealand. In addition they also prompted discussions with members of New Zealand Winegrowers about the consequences of the high incidence of GSyV-1 and whether more viruses should be tested in addition to GLRaV-3 as part of the evolution of the grafting standards.
Impact
To date, no adverse biological impacts of the fleck-like viruses (members of the family Tymoviridae that include GFkV, GRVFV, GAMaV, GSyV-1 and GRGV) have been reported, and it was even suggested by one international expert that these viruses could be added to GRSPaV as the “virome background” of a “healthy-looking” grapevine and excluded from sanitary measures. This statement clearly resonates with the results of this current survey. The ecological impact of these viruses is still unclear but according to their incidence, GRSPaV and GSyV-1 could inhabit the same ecological niche.
The detection of multiple species of vitivirus is more problematic than the fleck-like viruses as vitiviruses can be associated with diseases. However, due to their linked transmission, the management of GLRaV-3 is likely to also remove co-infecting vitiviruses from nurseries and vineyards resulting in only low vitivirus incidence; the results of the survey confirm the strong association between GLRaV-3 and the presence of a vitivirus under New Zealand conditions.
This survey confirms that besides GLRaV-3, there is a lack of, or at most very low, movement of the other ampeloviruses (GLRaV-1 and GLRaV-4) as they were only detected in very few plants and were not found outside the germplasm collection. In the case of GLRaV-1, the two positive plants were likely to have been propagated from the same Chardonnay Mendoza imported in 1971. In contrast, the vitiviruses were more common, especially GVA and GVG. These two viruses were detected outside the germplasm. These findings would suggest that under New Zealand environments the vitiviruses GVA and GVG are vectored.
The new sequencing technologies enable the understanding of the complete virome of an environment. That holistic view has changed our understanding of the place of viruses in the environment and challenge the automatic association of virus presence with disease. Historically, virus research has focused on the diseased plant and in grapevine viral-like symptoms are distinctive. This study highlights the presence of multiple viruses in healthy-looking plants.
To date, this is the largest survey of commercial grapevines using high throughput sequencing technologies and is larger than any other similar published virus survey for any plant host. This study sheds light on the virus diversity within New Zealand vineyards, the potential route of various viruses into New Zealand, and the projected impact of the viruses based on the current knowledge of their biology.
This article first appeared in the June / July 2018 issue of the New Zealand Winegrower magazine.

















